About DEFGermplasm
DEFGermplasm is built using the Python and Flask frameworks for backend development, while HTML, CSS, and JavaScript are used for frontend presentation and user interaction. The entire project is deployed using Nginx and Gunicorn.
DEFGermplasm. Available at: https://defgermplasm.com/
To report a mistake or problem on the DEFGermplasm platform, please submit a message through DEFGermplasm's Google Group. https://groups.google.com/g/defgermplasm
Data and Usage
The DEFGermplasm platform provides genome sequences and annotations, along with protein and nucleic acid sequences. It includes transcriptome data from diverse tissues and phenotypic data for edible nut species like pecan and hickory, as well as wood-producing species such as Chinese fir and phoebe. Users can retrieve genotype data from each species page, while the related phenotypic data will be available for download soon.
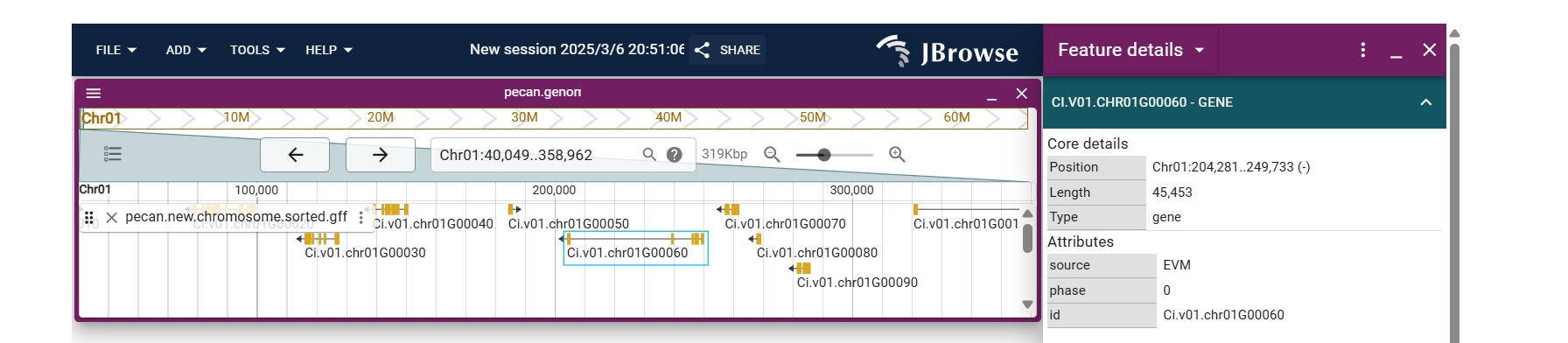
Users can access JBrowse from the species page or toolbar to explore gene locations and annotations within the genome.
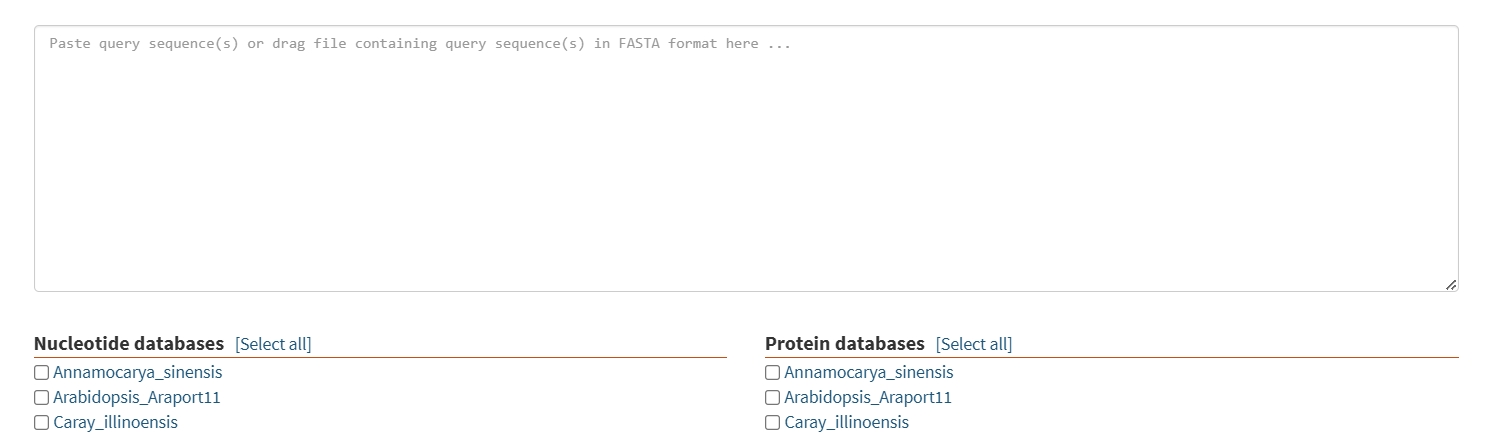
An online BLAST service where users can submit CDS or protein sequences for homology analysis. It includes a comparison library featuring 20 species such as walnut, pecan, Arabidopsis, rice, and poplar, enabling comprehensive sequence alignment.
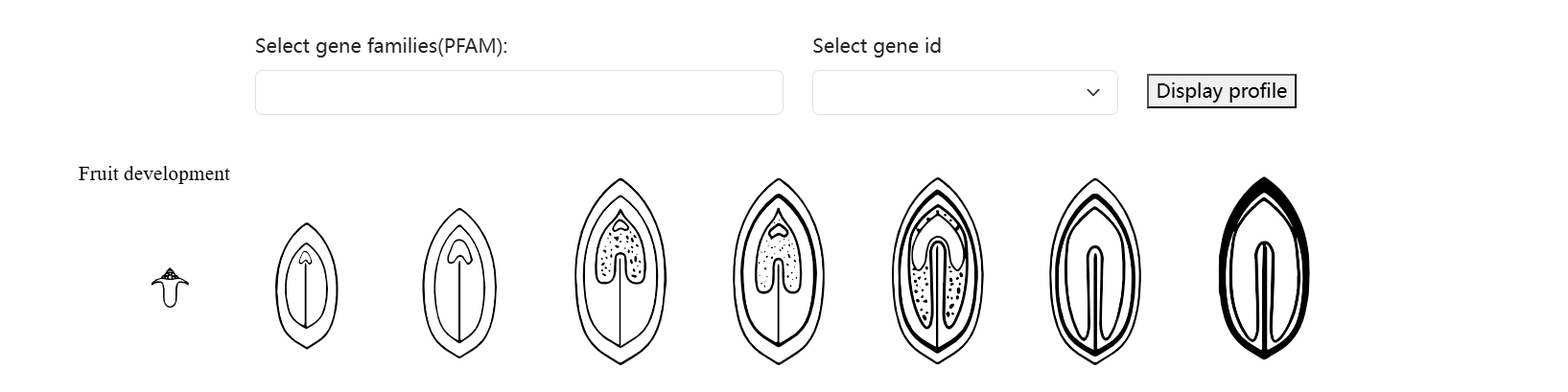
DEFGermplasm integrates transcriptome expression data with tissue locations, providing users with a visual representation of gene expression. Users can search for genes of interest by entering gene IDs or Pfam annotations to explore their expression patterns across different tissues.
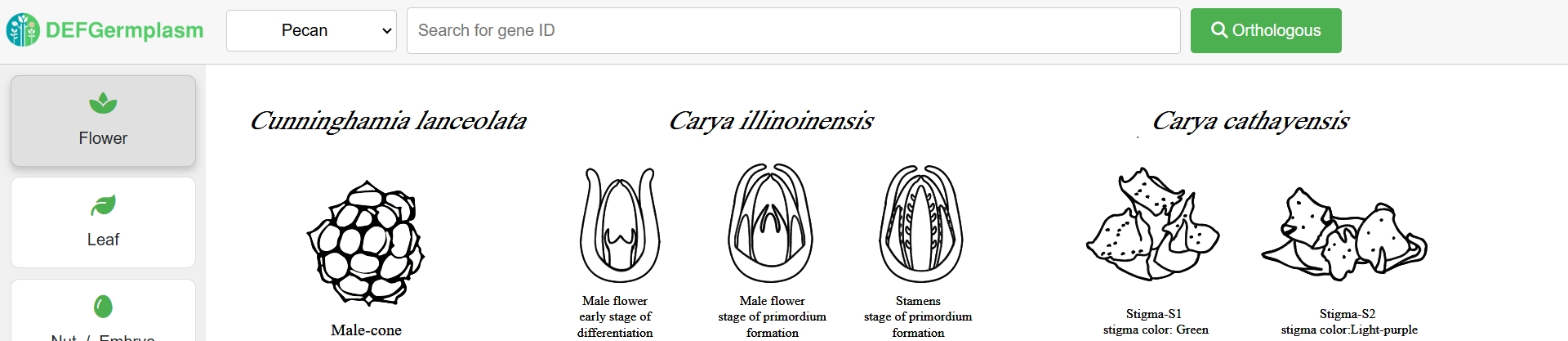
* Cross-species Expression Atlas
Orthologous genes from six species were identified using OrthoFinder. Users can search for orthologous genes through the search box and download gene trees and related data to facilitate their research. Additionally, users can explore the expression patterns of genes of interest across different species, allowing them to study both shared and species-specific gene expression.
You can download the data available on DEFGermplasm through the Download portal.